Highlights (en)
The article of the month: Best research paper award
The research paper “A Novel Indexing Method for Spatial-Keyword Range Queries”, co-authored by Associate Professor Christos Doulkeridis of the Dept. of Digital Systems, received the “Ralf H. Güting Best Research Paper Award” in the 17th International Symposium on Spatial and Temporal Databases (SSTD’21). SSTD is one of the flagship events of thespatial / spatiotemporal database community.
Presentation (video): https://www.youtube.com/watch?v=EkZDKP4KDXI
Multilevel study of Cutaneous Melanoma with Deep Learning Techniques
Ilias Maglogiannis
Professor
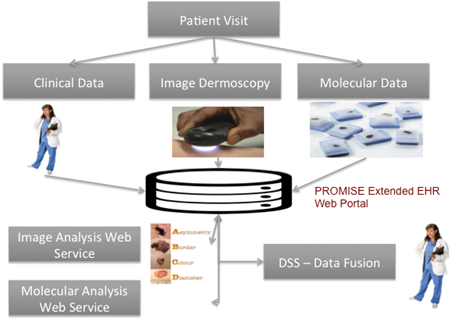
The study is based on multimodal analysis of demographic, clinical, imaging and molecular data
Image Dermoscopy data are utilized for the systematic monitoring of changes in skin lesions and the early diagnosis of Cutaneous Melanoma (CM).
An integrated Electronic Health Record SW is designed and developed, especially for the epidemiological management of a disease.
The Computational Biomedicine Laboratory (http://cbml.ds.unipi.gr/en/) in University of Piraeus implements in collaboration with other partners[1] the R&D project TRANSITION, aiming at the multilevel study of Cutaneous Melanoma (CM) through the intelligent combinatorial analysis of clinical, imaging and molecular data of patients with CM.
Cutaneous Melanoma with Deep Learning Techniques
CM is a malignant tumor of the skin’s melanocytes, the cells that are located in the basal layer of the epidermis and produce the pigment called melanin. Although CM accounts for less than 5% of skin cancer cases, it is responsible for the majority of deaths associated with it, due to its extremely aggressive nature. In recent decades, the frequency of CM incidents has been steadily increasing worldwide, due to aspects of the modern, western lifestyle (tourism and increased intermittent sun exposure, diet) and it is predicted that the increase will continue.
In the case of melanoma, the need for early diagnosis, much more intense than most types of cancer, becomes even more important because if the melanoma is detected early it can be successfully operated and cured completely. Otherwise, despite recent positive developments in the treatment of metastatic melanoma with targeted drugs and immunotherapy, the prognosis of melanoma at an advanced stage is particularly grim and the costs extremely high.
The project is extremely innovative from a technological point of view because:
- A multilevel molecular characterization of patients with melanoma is performed for the first time in Greece. More specifically, definition of potential inherited polymorphisms that may be associated with predisposition to the disease, detection of body mutations (mutations specific to cancer tissue) of each patient and study of the selected target genes’ expression of in cancer tissue will be performed.
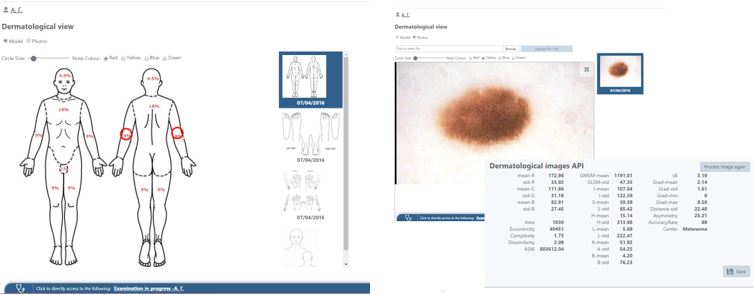
- Dermoscopy and confocal microscopy data are utilized for the systematic monitoring of changes in skin lesions and the early diagnosis of CM. More specifically, the correlation of the extracted visual characteristics with the molecular data will help in the designation of complex signatures for the diagnosis of CM and the understanding of the disease’s mechanisms. On the other hand, it will provide the basis for the extraction of new, improved and non-invasive disease classifiers through the utilization of new pattern recognition and deep machine learning techniques (Deep Neural Networks).
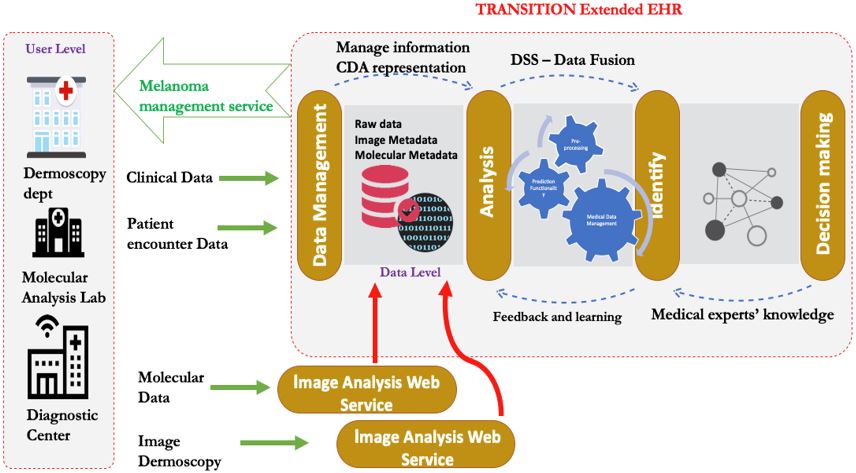
- An Integrated Electronic Health Record software is designed and developed, treating the CM as a realistic complexity model for the epidemiological management of a disease, completing all the individual subsystems and facilitating the processes of rational planning and medical decision-making support.
Although the software for the processing of dermoscopic images, developed by the research team in the Computational Biomedicine Laboratory, can work as a standalone application, it follows the principles of distributed computing and it is available as a programming interface (Application Programming Interface – API) to be integrated in the information system that is developed by the project. Its functionality is provided through Internet services (REST Web Services). REST (Representational State Transfer) provides a flexible architecture, allowing the exchange of different message formats for the communication between different information systems. Examples of these applications can be found at http://cbml.ds.unipi.gr/en/demos/
[1] The Transition project “Translating the diagnostic complexity of melanoma into rational therapeutic stratification” is co‐financed by the European Union and Greek national funds through the Operational Program Competitiveness, Entrepreneurship and Innovation, under the call RESEARCH – CREATE – INNOVATE. Datamed Systems Integration SA, the National Hellenic Research Foundation and the University Clinic of Cutaneous and Venereal Diseases at ANDREAS SYGGROS HOSPITAL participate in the project together with the University of Piraeus.
3D Modeling of UAV Networks (Unmanned Aerial Vehicles – UAVs)
Athanasios Kanatas
Professor
UAV networks have been proposed as a component of the next-generation wireless communication networks (beyond 5G).
Researchers from the Department of Digital Systems propose for a first time a simple yet realistic mathematical framework for the three-dimensional (3D) modeling of a UAV network consisting of a finite number of UAVs. The framework builds upon stochastic geometry tools.
3D MODELING OF UAV NETWORKS
UAV networks have been proposed as a component of the next-generation wireless communication networks (beyond 5G) due to specific advantages they provide in emergency situations, in large-scale concentration situations, even in military applications [1]. Their spatial modeling, however, remains a complicated task. Recently, the adoption of stochastic geometry tools proved capable of modeling extremely complicated frameworks of ground-based two-dimensional (2D) cellular networks [2]. Given the need for characterizing the interference inside a cellular network, stochastic geometry provides the potential to simplify the performance evaluation of such a network and extract some insights into their design.
3D Binomial Point Process has been proposed for the spatial modeling of UAV networks.
Until now, 2D Poisson point process has been widely used for the spatial modeling of users in current cellular networks. However, two major problems appear in this approach. The first one is that 2D modeling is not adequate for realistic UAV networks. Recently [3], researchers from the Department of Digital Systems indicated for the first time the necessity of studying 3D UAV networks. Specifically, it was shown both analytically and through simulations that current 2D models provide a more optimistic performance in terms of coverage, in contrast to a realistic 3D model. The second problem is that an aerial network cannot precisely be modeled through PPP, as different realizations of PPP constitute of different number of nodes. In fact, in realistic 3D aerial networks, the number of aerial nodes is finite. In such cases, a simple yet proper choice for the stochastic spatial modeling of aerial nodes, is the 3D binomial point process.
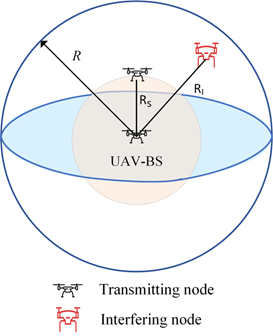
Within the context of the analysis performed, the 3D BPP was adopted for the spatial modeling and performance evaluation of a realistic UAV network, where a finite number of UAVs is deployed inside a sphere. A UAV – base station is deployed at the center of the sphere and assumed to communicate with the nearest UAV node. The link suffers from the presence of a dominant interferer. The model under investigation is depicted in the figure. The system was evaluated in terms of several performance metrics revealing useful insights about the design of UAV networks.
This paper conducted and published in the scientific journal IEEE Access by the Ph.D. candidate of the Department of Digital Systems Mr. Charalampos Armeniakos, under the supervision of Prof. Athanasios Kanatas, in collaboration with the Associate Professor of Univ. of Athens, Assistant Prof. Petros Bithas.
References:
[1] M. Mozaffari, W. Saad, M. Bennis, Y. Nam and M. Debbah, “A Tutorial on UAVs for Wireless Networks: Applications, Challenges, and Open Problems,” IEEE Commun. Surveys Tuts., vol. 21, no. 3, pp. 2334-2360, 2019.
[2] M. Haenggi, Stochastic Geometry for Wireless Networks. Cambridge, U.K.: Cambridge Univ. Press, 2012.
[3] C. K. Armeniakos, P. S. Bithas and A. G. Kanatas, “SIR Analysis in 3D UAV Networks: A Stochastic Geometry Approach,” in IEEE Access, doi:10.1109/ACCESS.2020.3036983.
Big Data Processing in NoSQL Systems
Christos Doulkeridis
Associate Professor
A novel method for unifying NoSQL storage systems for big data developers
Researchers from the Department of Digital Systems propose a novel method for unified data access to NoSQL stores.
Even though NoSQL stores (MongoDB, CouchDB, HBase, Cassandra, Redis, etc.) comprise the state-of-the-art technology for Big Data management, they still rely on different languages and programming APIs, thus hindering application development.
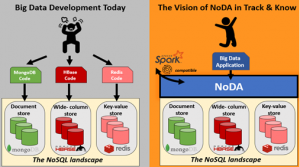
«Simple idea, yet very useful» [External reviewer]
In plain terms, NoDA is a programming API (see the blue layer at the right part of the figure) that consists of basic data access operators (such as filter, project, aggregate, sort), which are implemented for each NoSQL system separately, thus offering a simple and familiar language to the big data developer in order to implement applications, with the following advantages:
- It is simple to use and easy to learn, as it hides the peculiarities of each NoSQL system.
- It is unified, thus allowing code portability from one NoSQL system to the other, in the same spirit as in relational databases.
- It offers additionally a declarative, SQL-like interface, which makes it more user-friendly both for big data developers and data analysts.
This research work is carried out by PhD student Nikolaos Koutroumanis, supervised by Associate Professor Christos Doulkeridis.
More information:
- Paper: https://www.ds.unipi.gr/prof/cdoulk/papers/sstd19a.pdf
- Short video (teaser): https://www.youtube.com/watch?v=Rf6OYWP6lic
- Research project Track&Know: https://trackandknowproject.eu/
- General information on NoSQL systems: https://dl.acm.org/doi/10.1145/3158661